Chapter 1. Mirror Experiment Activity 13.1
Mirror Experiment Activity 13.1
The experiment described below explored the same concepts as the one described in Figure 13.1 in the textbook. Read the description of the experiment and answer the questions below the description to practice interpreting data and understanding experimental design.
Mirror Experiment activities practice skills described in the brief Experiment and Data Analysis Primers, which can be found by clicking on the “Resources” button on the upper right of your LaunchPad homepage. Certain questions in this activity draw on concepts described in the Statistics primer. Click on the “Key Terms” buttons to see definitions of terms used in the question, and click on the “Primer Section” button to pull up a relevant section from the primer.
Experiment
Background
As you have learned, researchers are able to sequence an organism’s genome by digesting or breaking it apart into smaller pieces, sequencing the resulting small genomic fragments, and then aligning these fragments based on sequence overlaps. Although it can be remarkably informative to look at the whole genome sequence of a single organism, researchers can also gain a great deal of information when they compare the whole genome sequences of two different species. They can identify mutations in the genetic sequences that are unique to one species or the other, and can isolate mutations that might be responsible for the different phenotypes observed in these organisms. How different are the sequences of the human and chimpanzee genomes? And if any differences exist between the whole genome sequences of humans and chimpanzees, can these differences help us understand how our species evolved?
Hypothesis
The Chimpanzee Sequencing and Analysis Consortium (CSAC) – a group of over 60 researchers at different institutions – hypothesized that by comparing the whole genome sequences of humans and chimpanzees, mutations that were unique to humans or to chimpanzees could be identified. They could effectively determine how different the chimpanzee and human genomes were from one another.
Experiment
Researchers isolated genetic material from the white blood cells of a chimpanzee. Using an approach similar to that outlined in Fig. 13.1, scientists constructed a preliminary whole genome sequence for this great ape. They compared the chimpanzee and human genomes, and counted the number of nucleotides that differed between these two sequences (Figure 1).
Photo credit: Anup Shah/Animals Animals-Earth Scenes
Results
As you will learn in later chapters, various types of mutations can occur in DNA sequences. Some of these mutations are a change in a single nucleotide, and are referred to as point mutations. Other mutations can involve larger chromosomal segments. A portion of a chromosome can be deleted and effectively removed from an organism’s genome. Chromosome segments can also be duplicated, resulting in extra material within the genome. Often, chromosomal mutations can affect thousands of nucleotides at a time.
The CSAC determined that, as a result of the combined action of point mutations and chromosomal insertions and deletions, the human and chimpanzee genomes differ in sequence by approximately 4%. Some of these differences between the human and chimpanzee genomes can be attributed to normal allelic differences between individuals in the same species. However, 1-2% of these differences are actually lineage-specific, meaning that they are not the result of normal variation within species, but true sequence differences that are unique to either the chimpanzee or human. Researchers were also able to identify certain chromosomal regions where mutations were more likely to occur, predict the rate at which certain genes diverged or evolved, and isolate a handful of mutations that could contribute to distinct human and chimpanzee phenotypes.
Source
The Chimpanzee Sequencing and Analysis Consortium. 2005. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature. 437, 69-87.
Question
As you learned in Fig. 13.1, to sequence a whole genome, researchers will typically break the genome into small fragments, sequence them, and then reassemble these fragments based on sequence overlaps. Two small genomic fragments that have overlapping sequences form a structure known as a contig. The CSAC relied on contigs to sequence the chimpanzee genome.
Question
jqnugG3m3287BXFhraQj5BaLi+eTUsimLjJvHjoDBmR01caMFqtQYNoF/4Hfqdkjv5YDAM3yLhNHIUt2xWgXdWJ1xgt3SRFF0j6RFK/n/FYmgRgq3oQqTItCkT6/6Tu56Mzs2Ks3X5/7epL6Vw4eDgC0otRMsQR06jHBXDxulI482UYTMWu+s753dC7tlIsumyrhWzwBV36o0+IDRqoKaXctghLKxjkmwTtaInTArByKm3y8NUf+a1PuPId855m1M9YabNExDtsKNGReYJJyV10SxI/zJu1mkSYBz6rbWCGLiGrn70KRCOQu1WU1368ZASBL+7hdU0fgee7HIYqKQKTNgCdoRIWtZaH57QOzQ3kfzcTqIAzJiVXdJVbG1Y7hP0ZcIAMI1QvP8tHREQsqYU3e1wNzYk97oIhIxCgUq1DHl7R3A4Kn0WBGN+GFl3xaWd73tUn7kHyXS6YuHsRaMHtaPcuh5yYeZE1TBHot9GhI9cOgw7t+4BktcNedpeP7eI0VpgqbOQtp6IZZMvG8+hxnn1a0fkn374/g0zKDn+QPaFb2JpW+Bglj7eQpoBbA6UXqN2H7nWaM5lJ7qD3ygfaQvN+ojWcPrSnuj/8D+VIxJGnkZQiZUn2En967lUatdfP/YXP/4pQxsqFcKMXkZS9VLB2Z/jXGxgO6xWlTYu62/JC/rJbNkTfC7Jf/7Qmn5oRnqlq6XARG01ck6dBqRtrMOGFzkGGoX9HZcRybx/2K5gU5jeK8yIBIvoq70AjVogPQoY6lbKg4QfSkPRiz0CgjfMZgyzkZNGS1KH8aL1SLAARH3vk3DV7PeUKk8gBrNo69p0KhZlOQmDgTsLgss8if7jLpksbP1brIe9FanTTQLC/v0NZJcYZIspWYC+wNbUyGpKo0+GEOUuk6H8Esn2TNnyRIVON3YfHqEFLmwL8V8EgS4fDECqVVbdq0p5jdXDFeXklyt3aqSFk8ayD78NKKFfLWNVLEQcy1aQTUZobQwQzuU6miz3z6WKtwZfGqvnKDODOcE3lQlT+mrnv6CK4Qovvm27Icnr4UVbminVGbPykbGJHMN9dWx85KUGcMA4WrsNNRYqoc3cP0/Ve6DwopGFAk2tbDZm+VPsg0xlADkFviTA0044U5nvUKonNeWYHgEvdQuQJWG0aue/z/cYSr8D1EeFutFEeo6PcdqM4a5+/jng9LqkyEZLVjskxPR0J1wfGhPkBVrflhg2oDWScB3+G4m6BCSEgU5Egsi8kfyCIOeqpxEp1BFKC9PPGQo2gJGD+DibEaRidzCdU8FdgeIndqIN3ZjCrU3skux6JWVXk9YT+Z4v0PDm19adWX+b3SFUnPiJvO/Ya2itzUEuA3gw+EksgZwWZ51Ugki509y9D6UU71AtGCyiVLUWGM7buw2j8rOlvTY0f6u0KzHQUDpfI/Rgd6fDGJDZ3E/ZelFepM7SZSE8G4MBo6OhxdpaXtxVyDIBjDT8tIICXm/8B2wpz4FFlctSYGeEBWHD2/lTyH5EddC9APwfSea3GaTkEQToiNIySF8qT9vTQD2SF3V/+C79WcM48AjqQkUNn0fAJBgkdhAgmYjPqhhufTnqp5yNHQwSpEJyQv7MoOwoyqo/SMHNpT4InL9rWg4KFFP2KVtg4HHlv/h8T2b7I1Ndst+Sc4bVrfv5IrxFDXUook4DhX3ZHcYBiBpBUqmxCeqOuWTbg/YTUAfIZ4tx6O8nPBYrXQd8c1ByDzMLO+lD3WBS2oVLUoA+My5zzReonky5YVVyBFk2AxB+z31ZfnpcVsSC8KuudvAk6+RIepLEzoDqv7aRMvHkSm6yVUYbRYJN8joM/MGQX+wWvvSKf6uBoPPEV8chDoTQSk98cxS+Pjk0PVOCQQQAk2EfM5gbYxM3f8rkp2LLnYVkezKvNLbokbewTCp2S0a8hVfwwDjNJKzv/2dOVRVAvasIpsNtPQj9GJJG5BAbfhIPYXsicSrvnymYPSsvb8xmEWuED7jVvI/YWbrP6SefXGoZ/az1Hv+A31kqjtLzlREjBXR+d9A3TT+Z+14opClFkY/2x+k56ZOSzNoT6N0NqdLkegwmLki2GQC7bliqD06Ace7A7WqUuvs0iyYI1GEq1ZqtvxgG8yJFP4GetdLhEdL3AgYRGFp7flB3lAtl6EZIsqCwbz+Hnyrp4PIWwrglVGMn2wvOk0Ld28Jw8H6wFrSfRQq/S8Kb/qyU2fBUIg3fge8mFTNI4uHqHi4E7n3nl/oifHLg9Wk//cQQJbaDVJDFMGhx5sm7WZPa5maz0UzQd6rYeOKXW0AarNuBQYeUYsDs7227D/4tuR+Nxn69N/fs70ozJTxeEAT+dZylsUbnuxJgWz9Z+3h+BUnGNMpedAeih9bFprUkQpxxmelBQ886OPLwthwenpG/tAA/toc7lFd0kw/DQsl8RlzGqo9+otxurX1TTbi9UsD+k16KectdrZAAhUJTcBpMawTBUdAkWsNIlIH7tOyttyhfFnGDf+NJrwfWF88P/nx0+fD26QTSgp+GQo4dLnr4S3iMPPRE88egIAiwNjoDdzYXVniX4HNuzLMmkm23/j9JPqj+AEDU4oncxN0twkqGyMwimW4NBCd6cIA/iIt3qsAWK9d6hzr/JrxxsFyZZkNwVccK3bsOZB9DjJtFEa71LpFeUp3DmLJBDDHXZU5+7+JheFBB4kDdMgAbSCLwlCrhNJDTmxqg7bX2XlWTqTm5wbVdDhThV3wP9EC6uSPjHklrVb3bLn7UoODpdWWs+tUmI0u0VFbV2ie08VZYksMpYdZ8jNRjPzYdi3Z0cd76pegpKdsmxlCMgHNMEdbDtBBahkG/EPxARc69lHMJ9mS+gyGthpACShvdM0IIEaX4SxUR1BcbtRKSjFT6rzeS/GMTo9iVX1+2qxDic1F30N3qTKSv8e0e60+pUfl6BH44+DymfzRGXwaBXxCFW6jzrpcjlPz9yaWFNw59ZQsfjGFRMGnYhtfpEZZFhCkavDQw94YI1/Yr5zinv4Gw==Question
SeBuQvsov0nIkTZSKu24DYKnFSaq4FFnAEPAgKIbU6IgNhuDmBbXtLX6z96YjUzr78kIM+RUsNyekGOf4UK2rPGotsh2MDrhAgi0xLKWgniP9O+VOcX8y2jxeGUk2qwgFZko24eaamN4NKln8tLmO74kiDerjG2WHsHCu/nUEcmXVQYT626Lhdt2ng8opzXAnH7sTts1wzAWQa5qYKDg2vgAGDqoO1FpleGNhFWJ1P6ptTcNphvMrRML1vj8SBT2IyqlTZR4igJtt16fBv9MNpGvlvfJO/YzkQFB3M3STId6RJL1ysT0FWOT+Dsxtbm0ID86ZFeqTtDZhHWfrTn5a+ETPWA+sjCegBW9NaW9NZz+ynE5lxxlP0XgH8vDXdfGU7oBlfdoCRTmypWTYl1a6NMgMaGLm5s+K1dmnuSKqeF/6QJZQM1h3KhPT7pEA3IQrMU+KJYv/sUIGHshpLTo3KuKZduTqpcOTvp75ERjcK98MJT6CbA8jTE3xK7Rpyw/P3e9yCeh/M8pMKqyptJIxcrle09sy+vy3wEIt5j1fukt5SgZjyz3b/Yhje7v5Ns+3hfa6FJNDmR2XqyYbENR2/R9H2FV6oxeyGfIMmZB5MV5w1okORWJwNzzFt5mxL0BVHNuAC6IpHipMbsICa1TX9Pfx4DQoprVnFx+bQLelqRp91MD65KhfGisqsfPtQtE6QrEPUHfyw+gq3FPVtO5r547QQh05KvNwQM3gT3poJIqATSKeTXHSj1Rsy5gGqmSCnZzewEwvOe1vQ/De7DHMa4IoBOGBEpFpHP3xFtTgLRRe+tXrew27x7s1Wi52JVQFLjvkOwmSIKveeNf6Gh1/xD+U/LTRH5/AMN7YwnfhZcW8s0oY7l6G2/bD7UVczYpJr7CRY7qqm070k0A7hfTGD8fAw3dkxWDxoVhhRh09dwPVbbAqwK7Ao3hb4qEozGHx9+Er20L01RZpavViLBTnH4igSiCuQ2CrH+xl3yKN0Gcks0Q44XnK3Cwozb45UBpb/wrqDT/zZa5Hix7ADLv8e5dWUQ05BaEcC+iImhWj3RogDAVA0DIU3esvEGMxgr0Ct/cAm/bZFimT6shUFcfBjdQykWGh8tqqrmXJmBz9iah9A7XypRxGocuUFlXBXWtjoipaHvs8SDzqsaTi9I5mkWQSmlKk1QQIwywWn90HD89+CrnDfFwGYk15vQdyHagh2nmpCKTjQT74R8Tgupj1cYYUuZOTvLxJJhibnLjDA+5LtWokSR21udmWAt+xvBgfQ/20GCh52suH4jyPMYpMSX/sWPEqf1TwAEdZBB4C1i4QxZ2jq4lV0EDvRqs1eDWNUMbbFTLvJdHEHFuu5fguxFNksYRZV20RugJvQuicw9rWLTxqa6ta/NDRUj0nH1WIO/ScPM6AAVyIil6rrTizNnxtKCFRgpgcBbA5AO9iOefqW9ILfeRwPHOq1kQfwBE1mJV6fzcmlRMmp4QyEihIUEi73vkIUGWraU4xF8iEIkwDmPb9FWeK3jH92gOUzWf+wjAB4b2G+balTaCqb8gm2hgtm7Hg+4avzrtQuwe/FgSUZB0nfErHsRC+TghFJMs07NHMMgYCCDtlQo8607wa27MuSq7xkWd3UqrPXBTyv1xpjRmUkvwWN3x+nM9/4PyUKS3anmD+/fbEBIFj34Ja4swj4wMo/Fdx+UkTG3db2AnxWO7lXj1046A4fd+TpVej7yqWXZyl/J76Og5vDzfPbu/z84kuuCP5T28NxzcqN5W5o1jSIx5UwpW9Y7g/g2r/6S6mxWj2X1dR7cfr8T1R4dWGtsh6khXQD9BSN50eV2dFTNZK441anpJLrYQe1q5YxG+OzUy85qWWLek9Y+Okw+tu+erQ7PQGCaMBoqhami+EOAnkhFWnMhYNBS52JHwK7UOecsuecaOweF5ijpItjF/4+CEDcyCmud5BUcI+Ae80/tseCgdJja/dZM6yKIhBVhmcIdwjY3wJQjVbtQY4uWylSIClp/CvMuiIsNz+J9bdr1Rk3XwK95A8CdvkWNiGfpS5XjERICggQQS7YkNrRF1GYMr2dmhbqPi3qV4H3AKYz72QedJgLBBl0BM2Cos/AW96vAY6a4TU+lsMsujbgbCPyJxrcogtRJH1Z7zQk6p9AE3Du5Nd2CL+6MGqU7vodBE2F+Wdd4oP/62OB+fDyBaYKNmxDfIkghmYb1B2l4vOLm/NjLtPwuanKJT9Tniitfp3wShLQbJyhNYh8MgCPmrBdlV/RwEbigSlR2m3Nj0Oa/GUCwWfcgxvSHWFioP4OwYCsoak0qLcZGffd36h4YtQpHIZqiEMb3QZSUdPDssuBUvqBoon+VHbl5CZyhP1zFfAzQ5lyqYVgJaFS6wAKnnawO4YdpV/X+J1z9hzjYeOV6w6TuQOw1PsgRnK0ePId/+vZsLmgPXMz3CSuSOc/OnWKwFETdaoSaDGE0DgqGyX2s9e6gBhJMotCU14+9JixoUuC+R+UCT8jBCRTG93ByYiX+EHHiWvTGkxagG3Owvw9yauMEoeRyLBnUZNj66nAQyVRyvuCMpoJ8pEPOlp7ps3Jc6aW3+77mRSLHBzI18oQC8twKHG8BZG6WmpX5GtCMg7DmNoIbm7S+CFZLufr2HkgVT3c0VEdpQQAUeTiwO5rjZDDNmkrNtg9ogFPsyHX2fD8eprW2pQm6S2RgSv0APpf64+Vg4YRMjWQpb9clAVXIFsBsfuhClzsUIhpGnJmHauInsafRxUnMHseInxip0ixoT3TrhZTuOyMp9OcttVPMntHy5OsO1JzHsCNqYoLoZnU3EyI/aEaPeaqY55jTvKRc+Qo/m0fMK+iDB7O3CLYXo3QGXLzDey7RV30lCwhj3rZHVWVjuOsRD4TUqWAOj/ctzW3GVVe9c55r8QdT1cSO6vc4cLpoHvp9Qo/tn1rjO0V6+UngqNw3gSU712DbQRwn81Iu1wQ/MxJ9Ay3zM/ZQ0/hpqZUg60ic1GnnIiKAV3T/KcHWV7x5lBvKixOLpmMtuEB5fcpeMzhAekJ/Nypcf6Sip1hfJa0QN+vwb5JtHNVV/nqNbYNQJGtdYqF/1MgD0zBNlv5LjZrhxqUBKH6HgLKU9CgoNR6e4gWfMoqbc00ZC22OafGwAtNWwSaLYknP/URPfWW3TaBae8B0kgOIT9cfADpSkyQdETQtZLqXw7+cLKkLO6TcnLHRT8yNK28LelGNbf1InomTadgWPOl8/dMjKAP+ZCd1aeS5d4Tgtm5OeijwU/fJs+Kv6A/0BlvzaHOkIPFIyF+ZTY9BNRJsdtB3iBnnpiGL1FmJnAvoTlAxJdz/O9iy7mTzsTesm1/jUIMGgaGrUBCKmefLQhzbaE3X8ncEgOqn0J/WTn6jJ8LAkQNN9knGHIeOMmUED7WrPX+nV/vhuVZF2ezVFIVZfPIJfjq8EmU0TjBC4SF2zAKsz9p12JHjju1Hp0bQZRmhZM1shLHL6nlaAGX0rW2uOEZSKHSGBo/xFjm7QqA0RLh4Z5vds8ci+DYwBcuf0z95FW3ffR/BbZFrtLJ/0eOUyU9WAMMcM4qo7cIcSUHaHAW8urti01drfKHSF95mXPL2GHpbEfQ/xpCxStd0++zaFqRiiVwqYQkq86vGQxxCW4znm+c8jOUpyhxX6hVu37lNxbH4WZUioB0EjO5vb514f7U/Xx9n/tYeTIKXh0qzfv36jIiL6CbNyA5FiFhy9QWcW3trpOYp0TrA9M6DlXEzsBfYzr2khI4WEAxGAD6J3WCbvCxaUzy5RczUrC/rwbG7SIzoNSvgQkS1SpZXwWFhxEtsWr+xusVUnRpAS+aAyTY+kogdl9PW1KPme4/3bDVZqW6f2HxKYyqntMtc8VNS4ABpx71uhCvCApW+oWSpSsKDXLatIp8hWfRK6ib1CyAiEdRyimMhtFUo/zu0g+jUp+H70eA==Question
The sequences of human chromosomes 1, 2, 3, 4, etc. are very close to those of chimpanzee chromosomes 1, 2, 3, 4, etc. Researchers of the CSAC compared the sequences of corresponding human and chimpanzee chromosomes and counted the number of base pairs that differed between them. In this manner, scientists were able to determine the extent to which human and chimpanzee chromosomes differed – or diverged – from one another. If analogous human and chimpanzee chromosomes demonstrate a divergence value of 0.010, this means that the sequences of the chromosomes in these two species differ by about 1%. The higher the divergence value, the more nucleotide differences exist between the sequences.

Question
When comparing the sequences of human and chimpanzee chromosomes,
the CSAC evaluated the divergence value for entire chromosomes, and also
for different chromosomal regions. In fact, scientists noted that certain
chromosomal regions are more likely to experience mutations and diverge in both
humans and chimpanzees. These researchers collected the following data for human
and chimpanzee chromosome 1 (this is a rendering of the actual divergence graph
depicted in Figure 2 of the paper); a regression line, which represents the overall
divergence trend for all areas of chromosome 1, has been superimposed over this data.
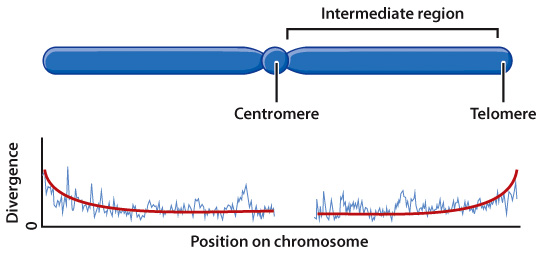
| Regression line | A line drawn on a scatterplot that depicts how, on average, the variable y changes as a function of the variable x across the whole set of data. |
Statistics
Correlation and Regression
Biologists often are also interested in the relation between two different measurements, such as height and weight or number of species on an island versus the size of the island. Such data are often depicted as a scatter plot (Figure 5), in which the magnitude of one variable is plotted along the x-axis and the other along the y-axis, each point representing one paired observation.
Figure 5A is the sort of data that would correspond to fingerprint ridge count (the number of raised skin ridges lying between two reference points in each fingerprint). While the data show some scatter, the overall trend is evident. There is a very strong association between the average fingerprint ridge count of parents and that of their offspring. The strength of association between two variables can be measured by the correlation coefficient, which theoretically ranges between +1 and –1. A correlation coefficient of +1 means a perfect positive relation (as one variable increases, the other increases proportionally), and a correlation coefficient of –1 implies a perfect negative relation (as one variable increases, the other decreases proportionally). Correlation coefficients of +1 or –1 are rarely observed in real data. In the case of fingerprint ridge count, the correlation coefficient is 0.9, which implies that the average fingerprint ridge count of offspring is almost (but not quite) equal to that of the parents. For a complex trait, this is a remarkably strong correlation.
Figure 5B represents data that would correspond to adult height. The data exhibit greater scatter than in Figure 5A; however, there is still a fairly strong resemblance between parents and offspring. The correlation coefficient in this case is 0.5. This value means that, on average, the offspring height is approximately halfway between that of the average of the parents and the average of the population as a whole.
The illustrations in Figure 5A and 5B also emphasize one limitation of the correlation coefficient. The correlation coefficient measures the strength of a straight-line (linear) relation. A nonlinear relation (one curving upward or downward) between two variables could be quite strong, but the data might still show a weak correlation.
Each of the straight lines in Figure 5 is a regression line or, more precisely, a regression line of y on x. Each line depicts how, on average, the variable y changes as a function of the variable x across the whole set of data. The slope of the line tells you how many units y changes, on average, for a unit change in x. A slope of +1 implies that a one-unit change in x results in a one-unit change in y, and a slope of 0 implies that the value of x has no effect on the value of y. The slope of a straight line relating values of y to those of x is known as the regression coefficient.
Question
9ar645nAknMqXj90fgVuE8PFyo7rOov+Ylnp0ImG+HvT8ik5wgt0PaFDkrqdM5ht7WfvbTJJdSjDqH6LjQ/WwjR5KW1F0nrmE2XwocYy6lk1/VNkwmtCUkbAzh/+x7yfDk0QqOWY2+Z3kqXmxXrNFX6/3oD6kJ7j+jyd7/rh/6gFuNIUtLm0bq6I9xEmRU/ZeRLfxWWbz5A1tF3MiecbTGucqkH0GHrDHtMPOsRr4RcPP0LI/HlAwxpt60qqJ6xFq3IXn55Kt8KxDHrU3AzZWszYE2GfEYbU91JDPBYBhg5OSH0aaoEab3wakggubaD8xGMBgKaMbW78hpnP7q2VU08hZHZrv6089xfQojxsEhTYr77qkX5G0ozVNOtK4vtIk1W2C768HgSi3fC0JM1xTySSGBb5don1OqMRS16JaEvZPT8SR3fZzsJTLtcEzsbxTVXdSRcXz8Unllm6y44CiP/nQ5i4B0gWnSfNuUcZwVZ9s2AucjjGLnCe/G87HAHz0jvk24OnIKlcruJCUkaedaBhpUJkqWUddjYOhqcclD57sZPp8nC/OWXV5u3ypluyZfycX6Au/eJr7xWMgYqwVRSaW8alNT3S0t4thHG8x6qvhpmywIAbV6v7t9SRZp4gTSBzOJACSJG/xWcGewngeQL2Xe+fPOHJrXsWeBFWfr4yR93NaTsBHN0sYP5M8UVvyZcCDxZV+W25woLIQux2YpXzyDcEwT0me5MJ3oVfdv/gRjVvW9UxXp82C92IFVTu1512Hgcuqcte0WxMn3tgPAjfOcsWtu75u7YKb0PfIRGQ/eh4mPAW7f5w5O67Mg/HoiNqW5Mr0M0QFWmg9asuLT6cUCOkQrerskURLTD2OEz53SgYRjEQY6I93sm6rIVy0uki8tkca/KJgRgCAK2NM67OqDUJHMdeilNgIMrf4PnuP8JeG6t3DYb1odoCRiBpsAPwUNYlDYd95JhQGBEQuf8bzjLNdwyjAkT3WR8pM8ElmwVtIjUImFcshgWKlhCXYIdgQKH5nmtasH3LxGewiS0IGpZidO7k00tvtq31mgmWgiNj+Ebag3y70wVKAZwC8GT7PRvHT+nnsHQq/i6Hvr5nDTf16ILY4lZABfIGOVTu/Upd91SnepwC6NGGay+01yye2xGDdYNZcf4s6/MurJRGpRcZ+0sD5ycVPTJSu3D0uECsYbaml3USYnyxxIn97ntQ6lNc4hqDk0gBysDMkIFOxWeC0o0DhzpSLLZaQgywqYwStJEDA5Kyvj5EEt0EwMTtrr7tbDGPG+iGN+kYQoASBFvDxLO/JH06q6r6ARwOEEG46nbGnwf+BpGPUzSkJ1B0c1VZ5ftsEMan/WkPJKDh3ptsR/4sPC3GvP2ROBLpMFbE5rnjxS2bH3UmjAsBqiNGh4WIpvoI/M/r43yZY5XV15L1lAmeDwxEEruieEtCQecWg+eP+djHK+rn0eJ7xf0cYjlnWUPi6ClENwzbIoFVVpjrUtBTZpTiD2aruosdnsRhzWvNB2mY7uGdLD84e92skj2gbPZgKdKoKGPYjmbk+2qD4c/p8Z3PDUbOk6mDtEPiFfzSFSvEPl/s1Z7UNqlpZr6i5qoWlqkmgDtehiHvwfq/YnSfWT3YQDxrlnZBvT/Tku93ZE/g9zT5UnoyzrnaUPqpoYdNw9jGiR9KX7UoM2IvASgRuX0vTu6/x9OjZiFY5mZU4rmPtcbCvdy/sgTz9V1OADDVsv+MjYcFXfYkNsCVnrK48HtbhPcQAXJpQSOiwmx++Zk0pXo1appiBbV8nv5vQM2J67qZysunVUypgdbUMoxg8vXnl52O6OUUGjo9iuU2Nv5j/8HhEbbDVTHzcQMNzKcli9FxdABsmf3galExQlXrqIwUUEcPjER5oRLzCu0ngoCucJFrZNjCt0nlBMRbQ+M8oICPu5/OP7uaO5eZa8qolW2E4Q8I7H0bU2SqlZcByiUA/HhzgN47fMt6iKpMM/vHB+XSBS2HK6Ci1xWHiFWmo9yS8NyrKqeVKvpsVqSgant2DEC/KNkAnbZ5YJ5tQP0yNFhkzMa4v6xpM67M0btLKMY2IGXgh+rkmOs+N15emGKK5ZhE0BOCpdbeeBoiGu6j51NBxn+Jvj5yAHgL6z7yBgdT2KfK87eZcELAi2cBq9Km8eTnxEL8Ms4ebjFzVFiUVPAAjKE/45otFZz9pcPHQFbQzk34bGnLr2jYDEWM7bVGpObWIPdJL0QJ4p5OqWA4BimUvAa0wpPngq57u5H6JKqg3slI1ofh3j7ImQO8mvUD7wJf3z8BO76GUXtTUuVohFW3+Gyg9/sSpxsjlzzFGea7eILOE5hhHWMfBSdI1L/7OUYCT1gO6pIsig==Question
XIvtHzSQqPysE4rxeLwo1mjU/b6tP6DhtrZH1ZErWnREG4JNXLjjeBDm2A3K/2Omf6Y3MJmClgV7Ne8mZU60QwlCruyXCVuWYdaVbOseCm/RUpZfGJG/VOF6W3aA7r8Kcen+NomkSz/R6HSaODY2zYwYFjpc3xHWldKimUZQdS4gtWohqCWjeHYdh/+W86gqh7wYIa8sdZ+GmpsBkzO1zEwUxJFT5i6BHbJtR7pYSgq/AfqUSPjoAWhedYRQHF/Tq8buOoZdFRzpNjMDnBTr57qLjIHjR0PjUfbSADb6qahZzIO9xUb7TVUaVmzLlYeM05RImYRUUE+rcx7OITPH4MJ2KxOnxB6LRIAvG/o2LUxAWmcCZK5zd1mZ6W1OMMtw4LvpRDU31KeO7+AmEbAazmgg4NNuoeFu6Qp3CNnVgywqzgFz29mzfAZHzp1nkwgBzAU8Ci5qKMOES79YjxfnT4wlbgCu7XxYaLRhPC8XYLXPwNn/MRVSF8yhXJh29P5Z54TCvYYWCP+Ma8NgOSbXnK+f7cvq9OFN4azUnjYqbYtBruTTvEqUCMIFIgpqh0TMM0i9UiGCDRwyuCgWQIhtonuZCeYm2ch2pI5vAPrEjI4F2a8vfOPNbqmvuf+1R3DCGfOvg1rpPIxWz8+CkrE//8OUEOUWLtXUI5kBqs/Lk3E347gD4/ZNMpfAw7PA7O+s5fuGbXRvN4QXpUfAGvhLm1YnQIBpkhPNmB/ezRefCiK/DSxFDilO7AASUimP/uNiHAjIYFfNNX1BDvET13igmEGiDyI7t+IQDf2kDafq/feU8Ug5HbXkQPSmjRJgmgUSO06v36DmrLh8FpfZElHkXJyOLhGqiNwUqxIAUT/Gtk0D2Za+DX/jp4BoLh4VCxNh1NQ32PycRT3rdgCDxwp7m2lKXUe5b79WwBFz2r9EZpMCqDft6BiqMtvJXyYQcM29HU8iVWc7eCsj3AweLaVth/lDwbyWSQY7T7wEE82ZzzSZ853DbIZ9V9gGhFp2KGQ6Dy8yUXDQVGRWtLzhdwYipk9OGzCo0Y7Hr7EInd49Fop5JJSUiy2kpi4WslWLHHblVPKk4dNNnSnLgwiGr17JNSTi0Q773i9Jm/gKHx1B5aCks9VZgr84cp4chpZ39xPUi8hinV7ULBz3fBHaWF3mt++NBBh2NglvSrooe+6v4rpRbc+SV6Y7M13mo+6Rq31aajSLRbDtEh/gAYormY0UDYJ8jznOKxT174pRvLRGSjp85ZdfQpigdfCHZxG2mV3i4p4obTyNge8OEr0wDq36+dn3hzA317a+IDVVRixVRIIYtZUKqtD9IxxvsTuXYHevyRx5KasLY5xiRiY3aKbLMsRoQjY1uQKHNvQ3PLq3jl998fYZnBlG1XnihfKHp3nwrEFrLov0JnePa+ivJNlEOhWOloHLUhyeLEAgyIyQT3i7V17qDpb9yRzef3QMAtfi33AOa+xw6ds7sQLwbgZykBFBKzdQbDZ8vzSKDvX/nrL+XebdkfTenjZzBBUhkZxZwEbxs61IVASTFu/PXLW1MbZCS+Wfep+M8kWSQh4DbG7E0rL/w30SY+x9j1jmRFmPscpwY6LQzHOxkM/874fm2JHmL7j1sLrxfCqeMwXwjNbRqidbq85y0P6yeRZ+QloAIo1yOLQD6NwirAKJLR0oZluebyg+H0pIYaNYtuincj229m05d3hAXlW+rRtx1kGXn0EzKcMcHlYIRD8weDlnJwgwcq2opis6sYbWZCFVQVniMGtO9SpRXvQDnZ9XYTjtvAe7hTwUF4EP918MBHWMNYu96IjrT7ixyuDEgSGfGFq+YPhh3gjhupB2vPHpa5+dqennGT/8Y2ELOJo2wchkPUlIYf4uwEE7BuffuuVLRIfRUXh9zJUdeJsuq8tM+I7ZRWT66EETMhhVs+6nqpaZpst0GRikHJcgHrpScZAIh9bMI6LIcl1SASMOACjMWfyyOY1Pvolfs8xJF4xEfDzTWs/2Yd9zw/EfQVZ92p+xmY4DD9cZ7co2gLTZp2CwcNcc+gvp6WcBS0/oX6YpR4BeTjGfaciSV54XJsDOytmHBvAgQnu5QfNGLMeK5i8pGMYGKdZ/8ScPmSNHXRaeX876h9PoT53gdWcHIhWaUKKkfnLtncxdn33gF3ODjkD87LyV/PUyVbW4KL01eqt8IR5NriYgou4BO8PGAJqnRC2QLDTzJ26EvV072zjbFEwpTkpt7PU6w54/tarGeKLjVbbVVk27qnVqobjanK04BjG7wqj9emX9ACben6uAireL7+8GvSJmE2p2lwaw4ufPkxW3PB7Fy1ahjs/u5I2RnLsBW9+q7ws3mYD4oYrQwRopdt8jVpxDceI5c6XOKRbAoYdDtDx4/MyE4AB97i6ldrhYYhtBYh0AMiRrXbQy5I2cp0GAzXSX72ZPtouHtdC/1e0Yc1WxXQ50h72D56FzxMxOIGX3pOElC0Irz4WPUoWYHFQQTrcZqO/l9KhIOMxyQtbufIalAsgrhDW6LmbCzC/0Wx8T2CmctRYNL0YrAWb7E65PRWqqs3mq4PV/OTTYHkw0OH0P7PnCwohg+i+b5Gjlu+tPNGFD